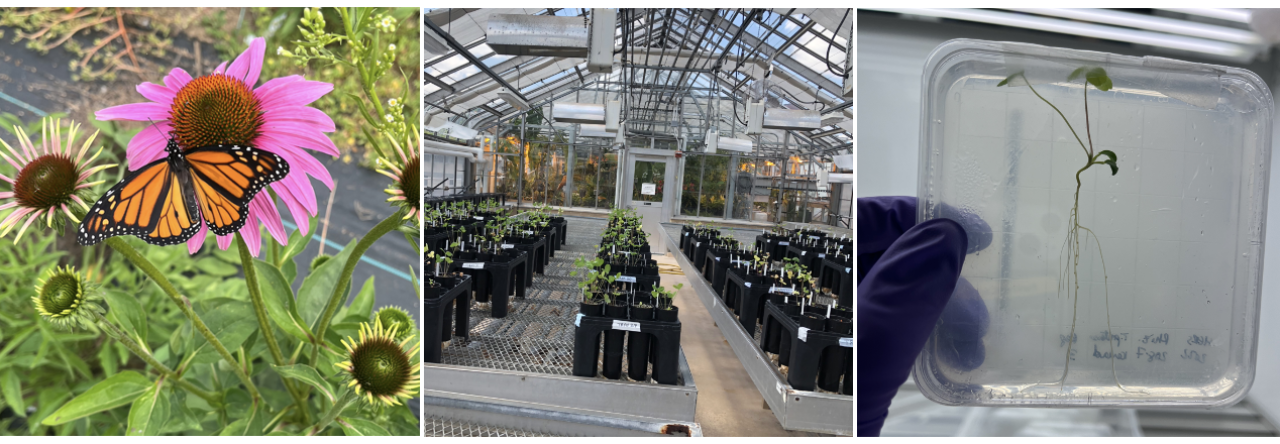
Each year, billions of pounds of pesticides are used to improve growing conditions for crops in the U.S., of which herbicides account for more than half of total usage rates. But what happens when herbicidal chemicals don’t stay in their intended area? How are plants and animals impacted when an herbicide infiltrates their habitats from neighboring agricultural fields? These are some of the questions I sought to address in the first chapter of my PhD thesis that was just published in the journal Annals of Botany. In particular, my research advisor, Dr. Tia-Lynn Ashman (University of Pittsburgh, Pittsburgh, PA), collaborator Dr. Regina Baucom (University of Michigan, Ann Arbor, MI), and I were interested in knowing how different wildflower species that typically live in the margins of agricultural areas respond to pollution from the herbicide dicamba, which has risen in use exponentially over the past several years and has been linked to numerous cases of ‘drift’ pollution, which occurs when herbicidal particles are taken up by the wind and carried to non-target areas at low concentrations. We focused on wildflowers because they make up diverse plant communities that are important food sources for a variety of animals, especially pollinators like bees. Therefore, by characterizing the effects of drift pollution on wildflowers, we also aimed to make predictions about how pollinators are indirectly impacted.
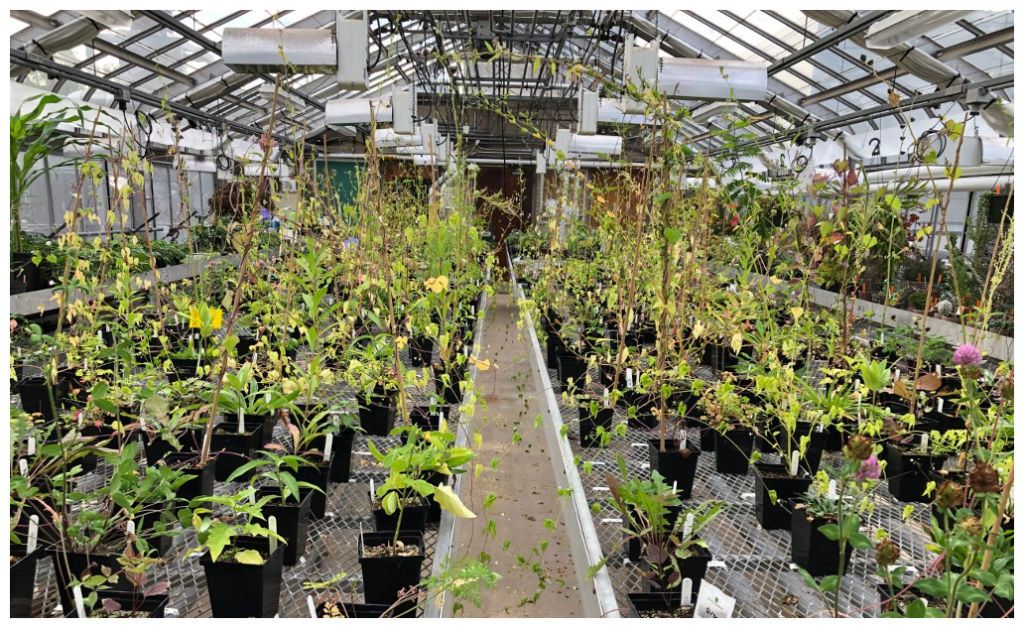

To address our research questions, we created two ‘synthetic’ communities of wildflowers in the greenhouse at the University of Pittsburgh, each comprised of 25 species that we previously identified as common inhabitants of agricultural margins in the U.S. We treated one community with an extremely small dose of dicamba to mimic what plants would be exposed to when drift pollution occurs and left the other community unexposed as a control. We tracked the growth and flowering of all plants by recording data on leaf and flower production and weighing samples of dry leaves, stems, and flowers to make size estimates. Using data on flowering, we constructed ‘networks of co-flowering interactions’ for the drift and control community, which show how frequently species overlap in flowering over time.

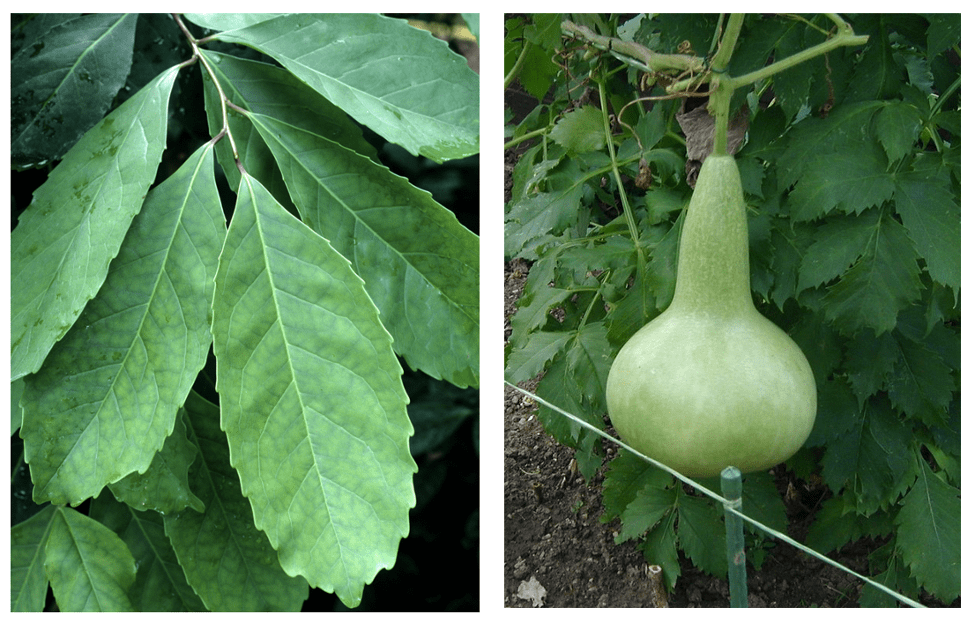
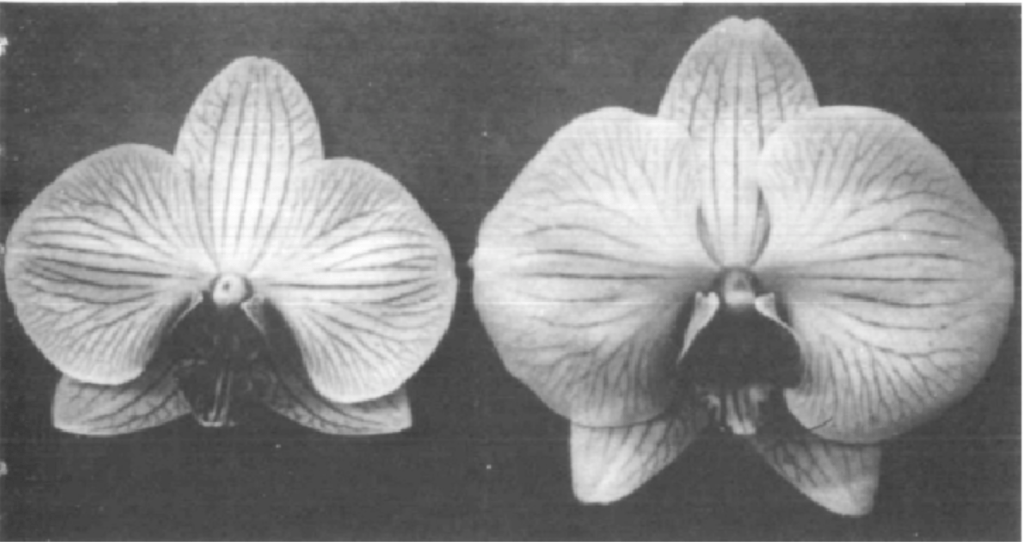
Through this experiment, we saw that there was considerable variation in how wildflower species responded to drift pollution. Many were sensitive and showed significant signs of damage immediately after drift treatment, such as stem twisting and leaf ‘cupping’ (when leaves fold inward to make a cup shape). A quarter of species were also negatively impacted by drift in terms of growth: plants of these species treated with drift were substantially smaller than their controls. Still, most species ultimately grew to comparable sizes regardless of the whether they were treated with drift or not, and a couple even grew to be significantly larger when treated with drift, possibly because the dicamba herbicide’s mode of action is to mimic the plant growth hormone auxin. The most interesting finding from this study, however, was how patterns of flowering changed. We found that most species were affected in terms of flowering time by drift—they either flowered significantly later or earlier when treated with drift than the control or flowered for shorter or longer periods of time throughout the duration of the six month-long experiment. Ultimately, this variation in flowering responses across wildflower species led to pronounced differences in the structure of coflowering networks between the control and drift community. In particular, the drift network was significantly less connected than the control, meaning that species tended to overlap in flowering less frequently. Additionally, groups of species that were more likely to overlap in flowering with each other were composed differently in the control network than the drift network. Altogether, these results suggest that herbicide drift has potential consequences for wildflowers in terms of changing which species are competing for pollinators at a given time, but also for pollinators in terms of shifting how many and which kinds of floral resources (since these are determined by which species are flowering) are available to them.

Overall, this study was important for showing how even very low levels of herbicide exposure can have profound effects on the growth and flowering of many wildflower species and also potentially the pollinators that would visit them. The next step we would need to better understand the consequences of drift pollution for wild plants and animals is to transition from ‘synthetic’ communities in the greenhouse to real ones in field conditions.